TDM 40200 Project 9 - Cross Validation
Project Objectives
Cross Validation is a fundamental machine learning technique used to evaluate the performance of a model. Instead of relying on a single split of the data, cross validation allows us to train and test on multiple subsets.
We will begin with data exploration and cleaning, and build a model using K-Nearest Neighbors, which will help us understand common issues and the motivation behind cross validation. The primary goal of this project is to understand why cross validation is necessary and why it is important; we aim to get familiar with the concepts and implementation, while also getting some introduction to different types of cross validation.
Dataset
-
/anvil/projects/tdm/data/nhanes/Nhanes_cvd_raw.csv
The dataset used in this project is the National Health and Nutrition Examination Survey (NHANES) data and it was gathered from the kaggle page.
This is an ongoing survey since 1999 conducted by Centers for Disease Control and Prevention with the aim to gain insights on American’s health and nutrition. It consists of interviews, health exams, and laboratory tests. You can learn more about it from CDC’s National Center for Health Statistics (NCHS) page.
|
If AI is used in any cases, such as for debugging, research, etc., we now require that you submit a link to the entire chat history. For example, if you used ChatGPT, there is an “Share” option in the conversation sidebar. Click on “Create Link” and please add the shareable link as a part of your citation. The project template in the Examples Book now has a “Link to AI Chat History” section; please have this included in all your projects. If you did not use any AI tools, you may write “None”. We allow using AI for learning purposes; however, all submitted materials (code, comments, and explanations) must all be your own work and in your own words. No content or ideas should be directly applied or copy pasted to your projects. Please refer to GenAI page in the example book. Failing to follow these guidelines is considered as academic dishonesty. |
Questions
Suppose we wanted to predict whether an individual has had a heart attack before. We would need a model that learns patterns and classify each patients. However, an important part of this is knowing how to evaluate the model. We will build a model using K-Nearest Neighbor (KNN), then explore and focus more on cross validation.
Question 1 - Data Exploration and Cleaning (2 points)
Before getting started, let’s understand what we are working with and what we need to prepare. We have done a lot of data exploration and cleaning throughout the Data Mine projects; we will follow a similar process here.
Please start by printing the head, shape, and columns of this dataset.
You will also almost immediately notice only from the head that there seem to be quite a few missing values. Let’s take a deeper look into that by counting the number of NaN in each column:
df = pd.read_csv("/anvil/projects/tdm/data/nhanes/Nhanes_cvd_raw.csv")
df.isna().sum()The output will indicate that the dataset has many missing data, which is not abnormal for a health and survey dataset especially where not all participants will complete every single exam, but we should handle it appropriately.
We will not just simply drop all rows with missing values using df.dropna() as we will lose most of the data. Instead, we will select a subset of relevant predictors, with our target being heart attack. Please use the following features.
features = ['Age', 'BMI', 'Systolic_BP', 'Diastolic_BP', 'Total_Colesterol', 'C_Reactive', 'Sodium', 'Saturated_Fat']The columns representing diseases, are coded in the NHANES dataset, seen at the table below:
| Code | Meaning |
|---|---|
1 |
Yes |
2 |
No |
7 |
Refused |
9 |
Don’t know |
. |
Missing |
|
You can see more information NHANES page, and related example in this decumentation. |
We can check that ourselves too.
for col in ["Heart_attack","Stroke","Angina","Coronary"]:
print(col)
print(df[col].value_counts())
print()Heart_attack
2.0 16553
1.0 421
9.0 66
Name: count, dtype: int64Above is an example output for Heart_attack only, and you will notice similar output with same coded values for other conditions as well. This confirms the NHANES coding scheme.
For us, values 7 and 9 are invalid so they will be removed.
df = df[df["Heart_attack"].isin([1,2])]Now we recode into a binary variable, such that 1 is for heart attack, and 0 is for no heart attack.
df["Heart_attack"] = df["Heart_attack"].map({1:1, 2:0})Now we will select the variables and drop missing values only for the used variables in this project.
df_clean = df[features + ["Heart_attack"]].dropna()1a. Print the shape, head, and columns of NHANES. How many rows and features are present? What are your other observations?
1b. Code and output that checks we only kept the rows where 'Heart_attack' column contains 1 or 2.
1c. Code and output that checks we converted to 1 and 0 for yes/no in heart attack.
1d. Head and shape of the new dataset.
Question 2 - KNN & Motivation (2 points)
This question will give a motivation into Cross Validaiton.
Now that we have the cleaned data, we will build a model using KNN. KNN is a simple algorithm that classifies points using the k-nearest training examples and a majority vote. The number of neighbors is defined with 'k'.
First, split features and target in the data by:
X = df_clean[features]
y = df_clean["Heart_attack"]KNN will calculate the distance between points and is sensitive to distance and scaling; without scaling, large scale variables will deominate calculations. StandardScaler shifts the data so the mean is 0 and standard deviation is 1, ensuring all features contribute equally. To do so, we need to import:
from sklearn.preprocessing import StandardScalerFor Train/Test split, import the following package:
from sklearn.model_selection import train_test_splitSplit the data into 80% train and 20% test using train_test_split after scaling features:
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y, test_size=0.2, random_state=42)-
test_size=0.2reserves 20% of the dataset for testing and remaining 80% for training. This is a very common split. -
Setting
random_state=42is for reproducibility, ensuring we get the same split every time the code is ran.
Now we are ready to train the KNN model. Scikit-learn library provides the KNN algorithm class. The second import will be used later to calculate the accuracy of classification models.
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_scoreknn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train, y_train)-
KNeighborsClassifier(n_neighbors=5)initializes the model with k=5 on the training set. -
knn.fit(X,y): The training data (X_train for us) is stored into X, and corresponding labels into y (y_train for us). Note that KNN does not learn a set of parameters during training like some other algorithms (e.g., linear regression), but rather stores the data points to calculate the distances later.
Now we want to measure how well the model worked.
train_predict = knn.predict(X_train)
train_accuracy = accuracy_score(y_train, train_predict)
test_predict = knn.predict(X_test)
test_accuracy = accuracy_score(y_test, test_predict)-
train_predictstores the generated predictions after taking in variables of the training set and calculating the predicted labels based on nearest neighbors. -
test_predictstores resulting predicted labels for the test set. The model makes predictions on the unseen set of input features (X_test) here. -
accuracy_score()takes in ground truth labels and predicted labels from the classifier. It measures (Correctly Predicted) / (Total Samples).
2a. All running code (given in question/examples) and outputs. Please include your own comments about the code and the processes.
2b. How does KNN work?
2c. Outputs of k=5 train and test accuracies. Explain what each values mean.
Question 3 - Problems & Overfitting and Underfitting (2 points)
With a very broad explanation, overfitting happens when a model learns the training data too closely and performs poorly on new data, while underfitting occurs when the model is too simple to capture the underlying pattern in the data. The following code chunk trains a KNN model for different values of k, computes accuracy on both training and test data, and stores the results to compare model performance.
# Store accuracy scores
train_acc = []
test_acc = []
k_vals = [1, 3, 5, 10, 20, 50]
for k in k_vals:
# Initialize KNN model with current k value
knn = '''YOUR CODE HERE'''
knn.fit(X_train, y_train)
# Predictions on data model has seen
train_predict = '''YOUR CODE HERE'''
# Prediction on unseen data
test_predict = '''YOUR CODE HERE'''
# Calculate the train and test accuracies and add to train_acc and test_acc
'''YOUR CODE HERE'''To get a better idea of what insight this provides us with, we can visualize this result as well. Please also write code to plot both training and test accuracy scores against k values in the same figure. You should see a plot like below as the output:
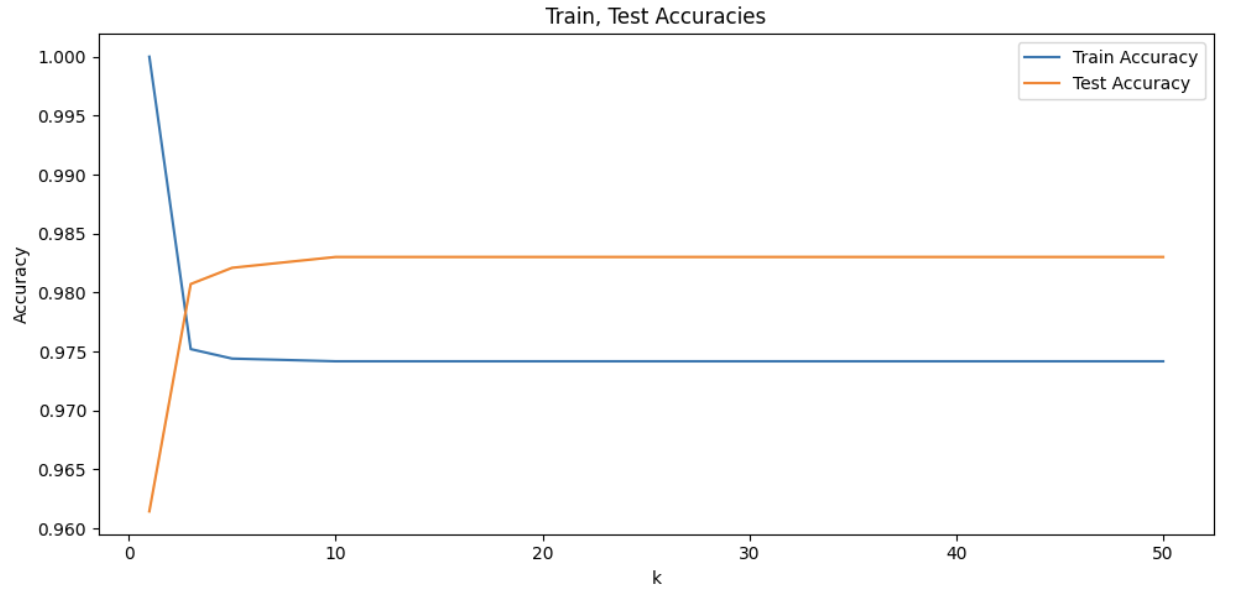
At the beginning of the graph, train accuracy is at 1.0, while being the same point where the test accuracy is the lowest. As k increases, train accuracy starts decreasing while testing accuracy starts increasing. With more neighbors, the model averages out data and noise better by being aware of broader patterns. Test accuracy seems to peak around k=10. As k continues to increase, we can notice the plateau; increase in number of neighbors is no longer effective in improving model’s accuracy and k values have become too large that it captures much of the global structure.
In other words, very small k values cause overfitting and large k values cause underfitting. Overfitting is when the model is very sensitive to local details and noise, and underfitting is when the model is too general and fails to capture specific patterns of the data.
|
We saw from the plot that the worst overfitting case occurred when |
The problem arises when, by chance, we obtain an unbalanced or unrepresentative train/test split. Depending on which rows we randomly land in validation set the accuracy can vary. A particularly easy or hard validation set would lead to error that does not accurately reflect true data.
We can illustrate this by training the same model multiple times on the same dataset.
scores = []
for i in range(20):
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y, test_size=0.2)
model = KNeighborsClassifier(n_neighbors=10)
model.fit(X_train, y_train)
predict = model.predict(X_test)
scores.append(accuracy_score(y_test, predict))
print(scores)The output is:
[0.9756769160165213, 0.9798072510325837, 0.976594768242313, 0.9811840293712712, 0.976594768242313, 0.9825608077099587, 0.9761358421294172, 0.9756769160165213, 0.976594768242313, 0.9784304726938963, 0.9733822854520422, 0.9752179899036255, 0.9770536943552088, 0.9779715465810005, 0.9761358421294172, 0.9775126204681046, 0.9710876548875631, 0.9788893988067922, 0.9798072510325837, 0.9761358421294172]The accuracy scores range differently although the model and data are the same. This single split approach gives us one outcome from a distribution of possible scores and we would not know which. Other important considerations is that we are also losing parts of data that could have been taken into consideration during training as 20% of the data is set aside, and we could end up with validation error that does not accurately reflect true data as an unlucky split can lead to a wrong selection.
This is where Cross Validation comes in.
3a. Code and output of training models for given varying k values and their accuracies.
3b. Plot both training and test accuracies vs k values. Please make sure all labels and titles are included.
3c. Write a few sentences on your observations and interpretations of the plot. As well, what do you observe as k increases, and at each extreme values of k?
3d. What problems does this cause when we try to make our predictions?
Question 4 - K-Fold Cross Validation (2 points)
K-Fold Cross Validation divides the data into K equal-sized folds. In each round, training is done on K-1 folds and the remaining fold is used for validation. The K validation scores are averaged to get a single estimate of the model’s performance.
We will continue to use Scikit-Learn. Scikit-Learn handles the entire cross validation process through cross_val_score function (see cross_val_score documentation for more details).
KFold class creates the indices to split the data into train and test sets. It splits the dataset into k partitions.
`KFold`class creates the indices to split the data into train and test sets. It splits the dataset into k partitions (see KFold documentation for more details).
from sklearn.model_selection import cross_val_score, KFoldBelow, we create KNN classifier and define our cross validation strategy. The dataset is split into 10 equal sized folds.
# Create KNN classifier using 10 neighbors
model = KNeighborsClassifier(n_neighbors=10)
# 10 Fold Cross Validation
kf = KFold(n_splits=10, shuffle=True, random_state=42)
cv_scores = cross_val_score(model, X_scaled, y, cv=kf)
# Print cross validation scores, mean accuracy, and the standard deviation.
'''YOUR CODE HERE'''-
In 'kf = KFold()', we set 'n_splits=10'. In the first round, 1st partition is the test set and remaining 2-10 are the train set. Similarly, in the second round, 2nd partition is the test set, and the 1st and 3-10 are the train set, and so on for all 10 rounds.
We also manually set 'shuffle=True' as KFold does not shuffle by default. This allows us to randomize row ordering. If there is no shuffling, the assignment of rows to folds would depend on their original positions; this can be problematic for sorted or grouped datasets, for example by age or dates, as it would lead to unrepresentative folds.
-
cross_val_score(model, X_scaled, y, cv=kf): We pass in the model, feature matrix, target labels, and the cross validation splitting strategy.
KFold from before just splits the indices. Training and evaluation of the model can not be done without knowing how to split, so we pass in the cv=kf argument.
Note that separating these components also provides flexibility, since cross_val_score() can accept other types of splitting strategy. For example, we can pass in StratifiedKFold or LeaveOneOut() (these will be presented in upcoming questions) instead of KFold.
Here, the model is trained 10 times and accuracy is evaluated for each fold. The function returns an array of scores corresponding to each cross-validation run.
4a. Output of all running code and the cross validation scores, mean, and standard deviation. Please add your own comments and documentation of process.
4b. Explain the role of n_splits and shuffle in KFold, and cross_val_score().
4c. What is your interpretation of the mean accuracy and the standard deviation?
4d. What do you think are advantages and disadvantages of k-fold cross validation?
Question 5 - More On Cross Validation (2 points)
K-Fold is standard, but there are other types of cross validation methods. Leave One Out Cross Validation (LOOCV) is a form of cross validation, where the number of folds is set equal to the total number of samples, n. Only one sample is used for validation per iteration and the remaining n-1 rows are used for training. This repeats until all rows have been used as the validation set once.
Just as an example, suppose we had a dataset with 5 observations. Then,
| Iteration | Training Set | Test Set |
|---|---|---|
1 |
Rows 2-5 |
Row 1 |
2 |
Rows 1, 3-5 |
Row 2 |
3 |
Rows 1,2,4,5 |
Row 3 |
4 |
Rows 1-3, 5 |
Row 4 |
5 |
Rows 1-4 |
Row 5 |
LeaveOneOut from Scikit-Learn provides the indices for splitting data into training and testing sets.
LeaveOneOut from Scikit-Learn provides the indices for splitting data into training and testing sets (see LeaveOneOut documentation for more details).
from sklearn.model_selection import LeaveOneOut|
LOOCV can be viewed as a special case of K-Fold CV. We would get the same result if we were to run |
Because the training set (size n-1) is almost the entire dataset (size n), we are generally able to obtain a low bias estimate with this method; the model is exposed to nearly all the data in each round. However, this also leads to LOOCV having high variance with the validation set being just a single point, thus very sensitive to outliers and noise.
At this point you might also realize LOOCV can have a high computational cost and is especially inefficient for large data, since the model needs to be trained once for every observation in the dataset.
print("Number of rows in dataset:", '''YOUR CODE HERE''')You should see:
Number of rows in dataset: 10892This implies fitting the model 10892 times is required if we are to use LOOCV.
Now, we will apply LOOCV (As previously mentioned, running this may take a couple minutes). Please complete the code below.
model = KNeighborsClassifier(n_neighbors=10)
# Initialize leave-one-out cross validation using LeaveOneOut()
loo = LeaveOneOut()
# Compute cross validation score
loo_score = cross_val_score(model, X_scaled, y, cv=loo)
# Print the total number of folds and mean accuracy
print("Number of folds:", '''YOUR CODE HERE''')
print("Mean Accuracy:", '''YOUR CODE HERE''')Note that we do not need to shuffle here since the concept of folds used in previous cases do not apply here; every single row gets used as test regardless of ordering. We do not see the problem of similar rows grouping into the same fold here.
Additionally, notice that we are just printing the mean and standard deviation of the accuracy. Comparing to regular K-Fold method, we were able to compute meaningful proportion using (Correct Predictions) / (Total in Fold); however, in LOOCV since each validation fold contains one row only, the model predicts either correct or incorrect. If we print the accuracy values, we will obtain an array of only 0s and 1s. You can see it for yourself too (try printing the accuracy value, and you will get: array([1., 1., 1., …, 1., 1., 1.]) ).
Mean value matters since it is (total correct predictions) / (total number of rows).
5a. Code to define KNN and perform LOOCV.
5b. Explain in your own words how LOOCV works, and how it differs from the previous method.
5c. What do you think are advantages and disadvantages of Leave One Out cross validation? What about in terms of bias and variance? When would you use LOOCV?
Question 6 (2 points)
Another version of cross validation is Stratified K-Fold Cross validation. You may recall our dataset is imbalanced, with only a small subset of individuals being a heart attack patient. Creating folds randomly may lead to some folds containing very few heart attack cases, resulting in unreliable evaluation results. Stratified K-Fold Cross validation is a way to resolve this type of issue by ensuring same class distribution as the overall dataset, making it particularly useful for classification problems (especially with uneven distributed labels).
`StratifiedKFold`from Scikit-Learn provides train/test indices for splitting data and keeps approximately the same percentage of target class samples as the original dataset (see StratifiedKFold page for more details).
from sklearn.model_selection import StratifiedKFoldmodel = KNeighborsClassifier(n_neighbors=10)
# Initialize Stratified K-Fold using StratifiedKFold() with 10 folds
stratified = '''YOUR CODE HERE'''
# Compute cross validation score
score = '''YOUR CODE HERE'''
# Print results, average accuracy, and mean accuracy
print("Stratified Cross Validation Scores:", '''YOUR CODE HERE''')
print("Mean Accuracy:", '''YOUR CODE HERE''')
print("Standard Deviation:", '''YOUR CODE HERE''')You will see output like:
Stratified Cross Validation Scores: [0.97522936 0.97522936 0.97612489 0.97612489 0.97612489 0.97612489
0.97612489 0.97612489 0.97612489 0.97612489]
Mean Accuracy: 0.9759457797322686
Standard Deviation: 0.0003582109670516864Each number in the score corresponds to the accuracy obtained in one fold. There are 10 since we used 10 folds.
Now, we will compare the class distribution inside each fold to verify that this method actually preserves it. For example, take regular K-Fold and our current Stratified K-Fold.
# Divide into 5 folds, shuffle, and we keep a fixed random seed
kf = KFold(n_splits=5, shuffle=True, random_state=42)
# Iterate through the folds
for i, (train_idx, test_idx) in enumerate(kf.split(X_scaled)):
label = y.iloc[test_idx]
# Print the fold number
print(f"\nFold {i+1}:")
# Count how many observations belong to each class in the fold
print(label.value_counts())-
kf.split(X_scaled)creates five pairs of index arrays, so for each fold it returns train_idx (indices of rows for training) and test_idx (indices of rows for testing). -
label = y.iloc[test_idx]: We use the validation indices to obtain the target label for the current fold. y is the target variable and test_idx has the row numbers belonging to the test set. We useilocto select rows by position index. -
Similar to before, "skf = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)" creates the object for stratified splitting. With our parameters, the dataset will be divided into five folds and data will be randomly shuffled before splitting.
-
Unlike regular KFold,
skf.split(X_scaled, y)requires both features and labels. In regular KFold, data splitting only worked based on position with no knowledge of the rows' class belonging, however in Stratified KFold, 'y' is required to separate rows into class group then build each fold with proportional sampling from each of them.
Output:
Fold 1:
Heart_attack
0 2142
1 37
Name: count, dtype: int64
Fold 2:
Heart_attack
0 2128
1 51
Name: count, dtype: int64
Fold 3:
Heart_attack
0 2127
1 51
Name: count, dtype: int64
Fold 4:
Heart_attack
0 2127
1 51
Name: count, dtype: int64
Fold 5:
Heart_attack
0 2106
1 72
Name: count, dtype: int64Please complete code below to see such for Stratified K-Fold as well.
# Stratified K-Fold
skf = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
# Iterate through the splits.
for i, (train_idx, test_idx) in enumerate(skf.split(X_scaled, y)):
# Get the test portion of the target variable for current fold
label = '''YOUR CODE HERE'''
# Print the fold number
'''YOUR CODE HERE'''
# Print the 0s and 1s count
'''YOUR CODE HERE'''Output:
Fold 1:
Heart_attack
0 2126
1 53
Name: count, dtype: int64
Fold 2:
Heart_attack
0 2126
1 53
Name: count, dtype: int64
Fold 3:
Heart_attack
0 2126
1 52
Name: count, dtype: int64
Fold 4:
Heart_attack
0 2126
1 52
Name: count, dtype: int64
Fold 5:
Heart_attack
0 2126
1 52
Name: count, dtype: int64Compare the outputs. In standard K-Fold, the heart attack cases range from 37 to 72 across folds. Model validated on Fold 1 has a set where heart attack patients are very underrepresented, thus the validation score would be unrealiable to be used as a good estimate. But Fold 5 had almost the double. Neither are accurate reflections.
However for stratified K-Fold, we can see that the number of heart attack cases is almost identical across folds. Each fold has about the same number of positive/negative cases.
6a. Code to define KNN and initialize Stratified K-Fold with 10 folds (make sure to shuffle).
6b. Output cross validation score, average accuracy, and mean accuracy.
6c. Code and output for class distribution for K-Fold Cross Validation and Stratified Cross Validation. Write a few sentences on your observation and interpretation.
6d. Explain in your own words how Stratified K-Fold works, and how it differs from other cross validation methods we have seen.
Submitting your Work
Once you have completed the questions, save your Jupyter notebook. You can then download the notebook and submit it to Gradescope.
-
firstname_lastname_project9.ipynb
|
It is necessary to document your work, with comments about each solution. All of your work needs to be your own work, with citations to any source that you used. Please make sure that your work is your own work, and that any outside sources (people, internet pages, generative AI, etc.) are cited properly in the project template. You must double check your Please take the time to double check your work. See submissions page for instructions on how to double check this. You will not receive full credit if your |